Project area A:
A1Protein/protein interactions of Ebola virus proteins as targets for new antiviral strategies
Stephan Becker
Prof. Dr. Stephan Becker
Institut für Virologie
Philipps-Universität Marburg
Hans-Meerwein-Str. 2
35043 Marburg
Phone: +49 (0)6421-28 66253
E-Mail: becker(at)staff.uni-marburg(dot)de
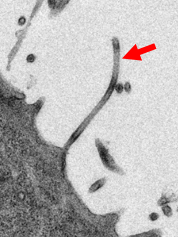
Electronmicroscopic picture of Ebola virus leaving an infected cell (red arrow). ©Schauflinger
Project description
The Ebola virus (EBOV) causes severe fever with extraordinarily high fatality rates. The matrix protein VP40 of EBOV plays key roles for the virus replication cycle and is regulated by homooligomerization. VP40 dimerization is crucial for the protein’s transport towards the plasma membrane where the dimers polymerize resulting in filament formation which enables virus budding. VP40 octamerization results in the down-regulation of viral RNA synthesis. Due to their central role as building blocks of the higher-order oligomers, dimers represent a promising target for antiviral intervention. A fragment-based approach was used to identify hits binding to VP40 crystals which will be developed into lead compounds in order to inhibit VP40 oligomerization.
Scientific goal:
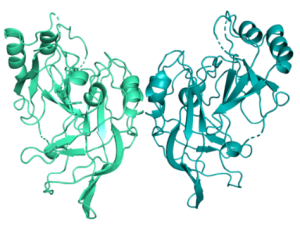
High-resolution crystal Structure of Ebola virus VP40 Dimers. ©Anke Werner
The project aims to develop lead molecules into antiviral compounds using structure-based drug design – a combination of protein crystallography with in silico methods – as well as cell culture experiments under BSL4- conditions for validation.
DRUID collaboration partners:
B1 Diederich/Kolb lab, A4 Heine/Reuter lab, D1 Steinmetzer lab, E3 Rahlfs/Przyborski lab
References A1: 1. *Hartlieb et al. (2007) PNAS 104: 624-9 2. *Hartlieb et al., (2003) J. Biol. Chem. 278: 41830-6 3. *Hoenen et al. (2005) J Virol. 79: 1898-905 4. *Möller et al. 79, 14876-86 (2005) J Virol. 5. Hoenen et al. (2010) J Virol 84: 7053-63. 6. *Gomis-Ruth et al. (2003) Structure 11: 423-33.
* own project-specific preliminary work
A2Development of eIF4A inhibitors as drug candidates and characterization of eIF4A-variants in Pathogens
Arnold Grünweller
Prof. Dr. Arnold Grünweller
Philipps-Universität Marburg
Bau C
Marbacher Weg 6
35032 Marburg
Tel.: +49 (0)6421-2825849
Fax: +49 (0)6421-28 25854
E-Mail: arnold.gruenweller(at)staff.uni-marburg(dot)de
Project description
The cellular RNA helicase eIF4A is an excellent target for the development of broad-spectrum antivirals. During initiation of viral protein synthesis, many viruses, especially corona viruses, rely on this enzyme, which can efficiently and specifically be inhibited by rocaglates. In project A2, we would like to further develop rocaglates for potential clinical trials by e.g. nebulizing these compounds for local application into the respiratory tract and by creating a detailed side effect profile. In addition, new eIF4A inhibitors will be screened and characterized. The systematic mutagenesis of known coronavirus sequences should allow a prediction of rocaglate sensitivity in newly emerging coronaviruses. Finally, the therapeutic relevance of rocaglates in different DRUID-relevant pathogens expressing eIF4A variants will be investigated.
![]()
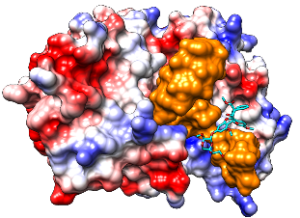

RNA clamping onto the surface of eIF4A by the rocaglate Silvestrol and evidence of inhibition of (corona)viral protein synthesis by the rocaglate CR-31-B (-).
Scientific goal:
Rocaglates will be further developed for their testing in clinical trials and the possibility to predict rocaglate sensitivity in emerging corona viruses and other DRUID-relevant pathogens will be evaluated.
DRUID Collaboration partners:
B2 Ziebuhr lab, B4 Grevelding lab, B5 Schlitzer lab, C6 Häberlein lab, D1 Friebertshäuser/Steinmetzer lab, D3 van Zandbergen lab, D4 Hermosilla/Mazurek/Taubert lab, E4 Spengler lab, E6 Schiffmann lab
References A2: 1.*Biedenkopf et al., (2017), Antiviral Res. 137: 76-81; 2. *Müller et al., (2018), Antiviral Res. 150:123-129; 3. *Elgner et al., (2018), Viruses. 10(4): 149; 4. *Glitscher et al., (2018), Viruses. 10(6): 301; 5. *Henß et al., (2018), Viruses. 10(11): 592; 6. *Müller et al., (2020), Antiviral Res. 175:104706; *7. *Müller et al., (2021), Antiviral Res. 186: 105012; 8. *Blum et al., (2020), J Cell Mol Med. 24(12): 6988-6999; [*own publications].
A3Posttranslational protein modifications as Achilles’ heel of pathogenic RNA viruses
Friedemann Weber
Prof. Dr. Friedemann Weber
Institut für Virologie
FB Veterinärmedizin
Justus-Liebig-Universität Gießen
Schubertstraße 81
35392 Gießen
Phone: +49 (0)641-99 38350
E-Mail: friedemann.weber(at)vetmed.uni-giessen(dot)de
Project description
Due to their small genome, viruses are highly dependent on functions of the host organism. Many of those functions are regulated by cell-encoded posttranslational protein modifications for which a substantial number of pharmaceutical inhibitors are available.
Rift Valley Fever Virus (RVFV) is a mosquito-borne zoonotic pathogen endemic in parts of Africa. In large and devastating outbreaks, it typically kills thousands of farm animals and hundreds of humans. In the preceding funding period, we used a high-throughput genetic screen and identified a pro-viral host cell factor for RVFV that binds to posttranslational protein modifications. Inhibition of this factor in a human organoid model reduced viral RNA synthesis and progeny particle production. In addition, proteomic analyses showed that a viral protein is modified is a manner that the host cell factor can bind, and mutation of the relevant site led to a reduction of viral RNA synthesis.
Scientific goal:
We aim to elucidate the molecular mechanism and exploit it to specifically inhibit RVFV infection. Moreover, we will test available pharmaceutical inhibitors, and also screen for other pathogenic RNA viruses that may depend on this mechanism.
DRUID Collaboration partners:
A1 Becker, A2 Grünweller, B2 Ziebuhr, C1 Bender/Hildt, D1 Friebertshäuser/Steinmetzer, E3 Rahlfs/ Przyborski, E4 Spengler, E6 Schiffmann, E7P Krijnse Locker
References A3: 1. Wuerth & Weber (2016) Viruses 8, 174*. 2. Barr, Weber, Schmaljohn (2020) Fields Virology, vol 1, p 706-749*
A4Search for lead structures to inhibit the chaperone IpgC from Shigella
Andreas Heine, Klaus Reuter
Prof. Dr. Andreas Heine
Institut für Pharmazeutische Chemie
Marbacher Weg 6
35032 Marburg
Tel.: +49 (0)6421-28 21313
Fax: +49 (0)6421-28 28994
E-Mail: heinea(at)staff.uni-marburg(dot)de
Prof. Dr. Klaus Reuter
Institut für Pharmazeutische Chemie
Philipps-Universität Marburg
Marbacher Weg 6
35032 Marburg
Tel.: +49 (0)6421-28 25845
Fax: +49 (0)6421-28 28994
E-Mail: reuterk(at)staff.uni-marburg(dot)de
Project description
Bacteria of the genus Shigella invade the epithelial cells of the colon, which results in the severe inflammation of the large intestine. Known as bacterial dysentery or Shigellosis, this causes a large number of deaths, foremost in developing countries. The Shigella specific chaperone IpgC interacts with numerous further pathogenicity factors and is prerequisite for the virulence of this organism. In the absence of a “substrate protein”, IpgC forms a homodimer, which is essential for its stability. We use IpgC as a target protein for the structure-based design of compounds against Shigellosis by preventing IpgC homodimer formation and/or binding to substrate proteins. By now, we have established a protocol which reproducibly yields excellently diffracting IpgC crystals. Using a fragment-based approach, we have identified a number of IpgC “binders”, some of which we were able to expand significantly. In addition to protein crystallography, we use “Microscale Thermophoresis”, Isothermal Titration Calorimetry” and a “Thermal Shift” assay to study the influence of such molecules on homodimer formation, on the ability to interact with “substrates” and on stability.
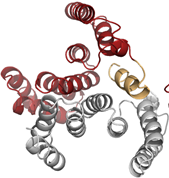
Crystal struc-ture of ho-modimeric IpgC. ©Klaus Reuter
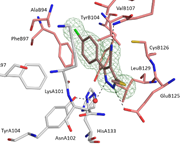
“Follow up” compound bound to IpgC. ©Marina Gardonyi
Scientific goal:
In addition to further structural information on IpgC, our main goal is the optimization of the compounds identified so far using them as lead structures in the development of anti-Shigellosis compounds.
DRUID Collaboration partners:
A1 Stephan Becker, B1 Wibke Diederich / Peter Kolb, B7 Franco Falcone, D1 Eva Friebertshäuser / Torsten Steinmetzer
References A4: [1] Agerberth et al. (2005) World Health Organ [2] Williams & Berkley (2018) Paediatr Int Child Health 38:50-65. [3] Sansonetti (2001) Am J Physiol Liver Physiol 280:319-323. [4] Parsot et al. (2003) Curr Opin Microbiol 6:7-14. [5] Lunelli et al. (2009) Proc Natl Acad Sci USA 106:9661-9666.
A6Targeting the highly divergent actin superfamily in malaria parasite transmission
Ross Douglas
Dr. Ross Douglas
Biomedical Research Center Seltersberg (BFS)
Molecular Infections Biology
Justus Liebig Universität Giessen
Schubertstrasse 81
35392 Gießen
Tel.: +49 (0)641-99 39145
Fax: +49 (0)641-99 39129
E-Mail: ross.g.douglas(at)ernaehrung.uni-giessen(dot)de
Project description
Malaria remains one of the most devastating diseases and is caused by single celled parasites called Plasmodium. These parasites are transmitted between people by Anopheles mosquitoes. The parasite needs a set of diverse proteins that enable it to transmit to the mosquito vector, including members of the highly divergent actin cytoskeleton and its regulators. The parasite cytoskeleton has unique properties in order to transmit and, given its essential nature in various parasite processes at different life cycle stages, contains promising targets for novel malaria therapies. We make use of target validation approaches to identify and characterize novel transmission blocking targets that could be used to control disease spread.
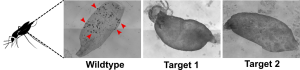
Target validation approach has identified novel transmission blocking targets.
Scientific goal:
We aim to characterize identified proteins of interest using a variety of in vitro and in vivo methods with a view to identify novel compounds that selectively target these proteins and thus serve as transmission blocking drug candidates.
DRUID Collaboration partners:
A7 Przyborski, B1 Diederich/Kolb, B7 P Falcone, E3 Rahlfs/Przyborski
References A6: [1] Douglas et al. (2018) PLOS Bio e2005345; [2] Douglas et al. (2018) Malaria J 17:3191898-905; [3] Douglas et al. (2015) Trends Parasitol 31(8):357-362.
A7Plasmodium chaperones, co-chaperones and their interactions as a target for drug development
Jude Przyborski
Prof. Dr. Jude Przyborski
Interdisziplinäres Forschungszentrum (iFZ)
Justus-Liebig-Universität Gießen
Heinrich-Buff-Ring 26-32
35392 Gießen
Tel.: +49 (0)641-99 39114
E-Mail: jude.przyborski(at)ernaehrung.uni-giessen(dot)de
Project description
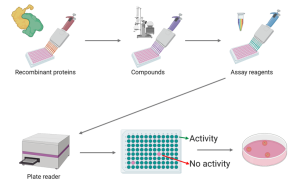
Overview of assay design
Malaria parasites invade and live within mature human red blood cells RBC). To enable their survival, the parasite renovates it’s chosen host cell to its own advantage. Infected red blood cells become sticky and adhere to the lining of small blood vessels, and also coat themselves with proteins which enable them to become invisible the immune system. This unfortunately causes disease in the patient, and eventually leads to death. We have recently identified a number of important molecular players which are essential for this renovation process, including members of the so-called HSP70 and HSP40
families. It is the goal of this project to block the function of HSP40 and HSP70. If we can do this, parasites are likely to be cleared from the bloodstream, relieving the severity of disease. To do this, we will establish a number of assays to measure the activity of HSP40/HSP70, and use these to search for compounds which reduce this interaction. Promising compounds will then be tested directly on parasites for their ability to reduce host cell modification.
References A7: 1. Diehl et al. (2021) PLoS Pathogens 17:e1009969 2. Zhang et al. (2017) Sci Rep 7: 42188 3. Charnaud et al. (2017) PLoS One 12: e0181656 4. Külzer et al. (2012) Cell Micro 14: 1784-95 5. Külzer et al. (2010) Cell Micro 12: 1398-1420







